BENCHMARKING RESULTS FOR RiPPMINER-GENOME
|
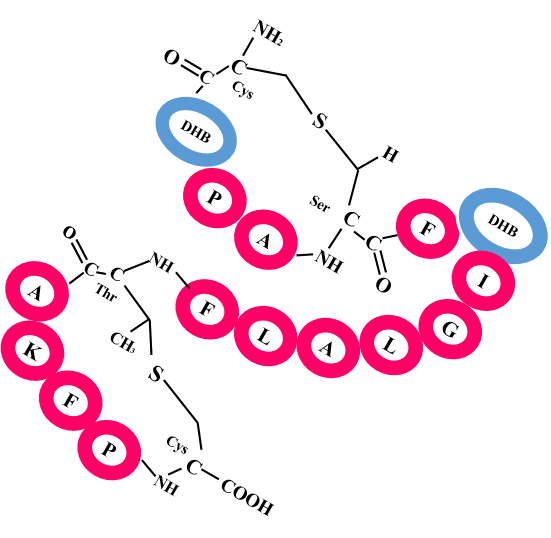
Correct identification of RiPP BGC/modifying enzymes and prediction of RiPP class could be done in 99% of cases. In more than 70% of cases the prediction corresponding to the correct precursor peptide, cleavage site and cross-link pattern was ranked among top three. |
| RiPP Class | BGC/GENOME Test Set | Correctly Predicted BGC | Correct Modification Ezyme Prediction | Correctly Predicted RiPP-Precursor Peptide | Correct Leader-Cleavage Prediction | Correct Cross-links Prediction |
|---|---|---|---|---|---|---|
| Lanthipeptide | 100 | 100 | 100 | 65 | 65 | 58 |
| Lassopeptide | 54 | 54 | 54 | 33 | 32 | 32 |
| Thiopeptide | 44 | 43 | 43 | 28 | 28 | 28 |
| Cyanobactin | 26 | 26 | 26 | 19 | 19 | 19 |
| LAPs | 9 | 9 | 9 | 4 | NA | NA |
| Glycocin | 12 | 12 | 12 | 3 | NA | NA |
| Head-To-Tail cyclized Peptide | 9 | 9 | 9 | 5 | NA | NA |
| Linardin | 5 | 5 | 5 | 1 | NA | NA |
| Bottromycin | 3 | 2 | 2 | 1 | NA | NA |
BENCHMARKING RESULTS FOR RiPP PRECURSOR PREDICTION FOR LANTHIPEPTIDE,
THIOPEPTIDE, LASSOPEPTIDE & CYANOBACTIN
BENCHMARKING RESULTS FOR LANTHIPEPTIDE MODIFIED RESIDUES PREDICTION
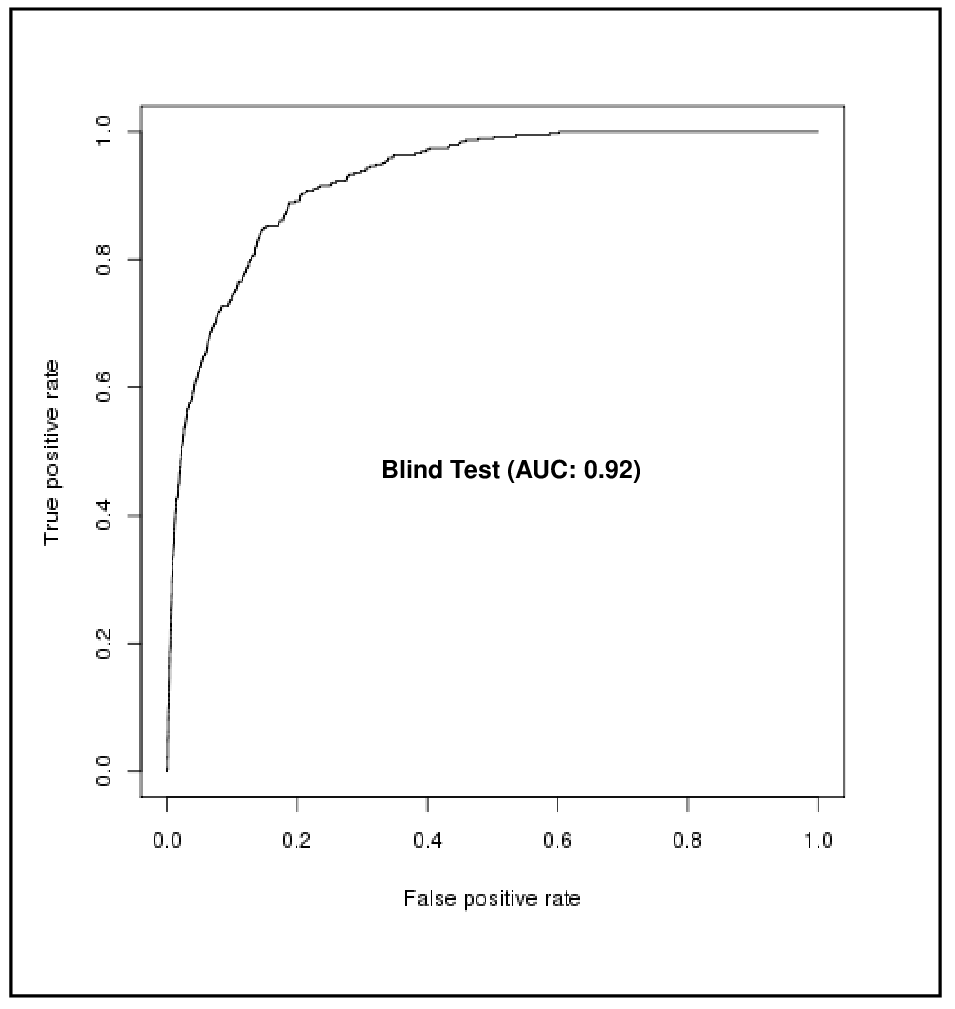
Method: Support Vector Machine (SVM)
| False Positive Rate (FPR %) | True Positive Rate (TPR %) |
| 10 | 78 |
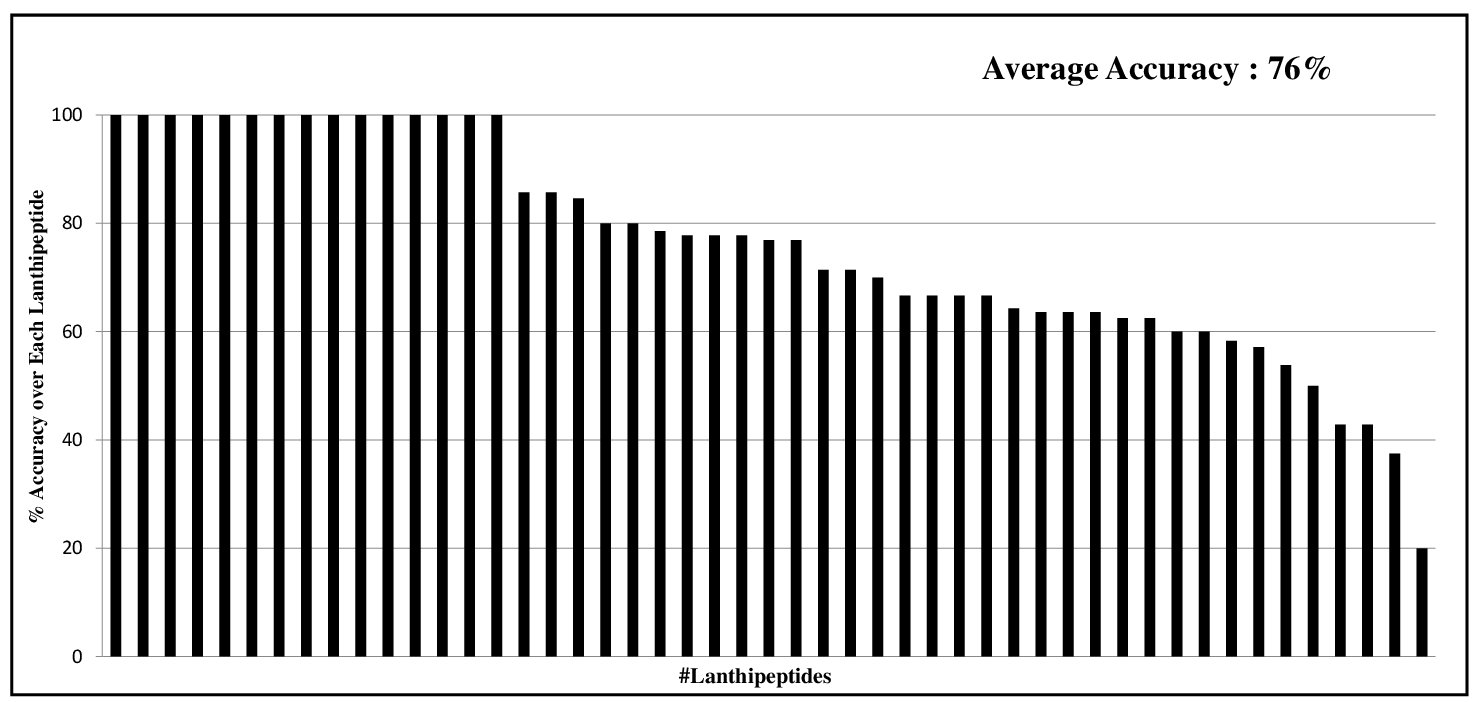
|
The Machine Learning (ML) Model was trained on 49 Lanthipeptides and was Tested on the remaining dataset of 49 Lanthipeptides in the Blind Test. The result shows here are the percent Accuracy for each of these 49 Lanthipeptides. The % Accuracy was calculated by counting the number of correctly predicted Modification States of Ser/Thr/Cys Residues by ML Model divided by total number of such residues present in the Core region of each Lanthipeptide. The average % Accuracy was 76%. |